正在加载图片...
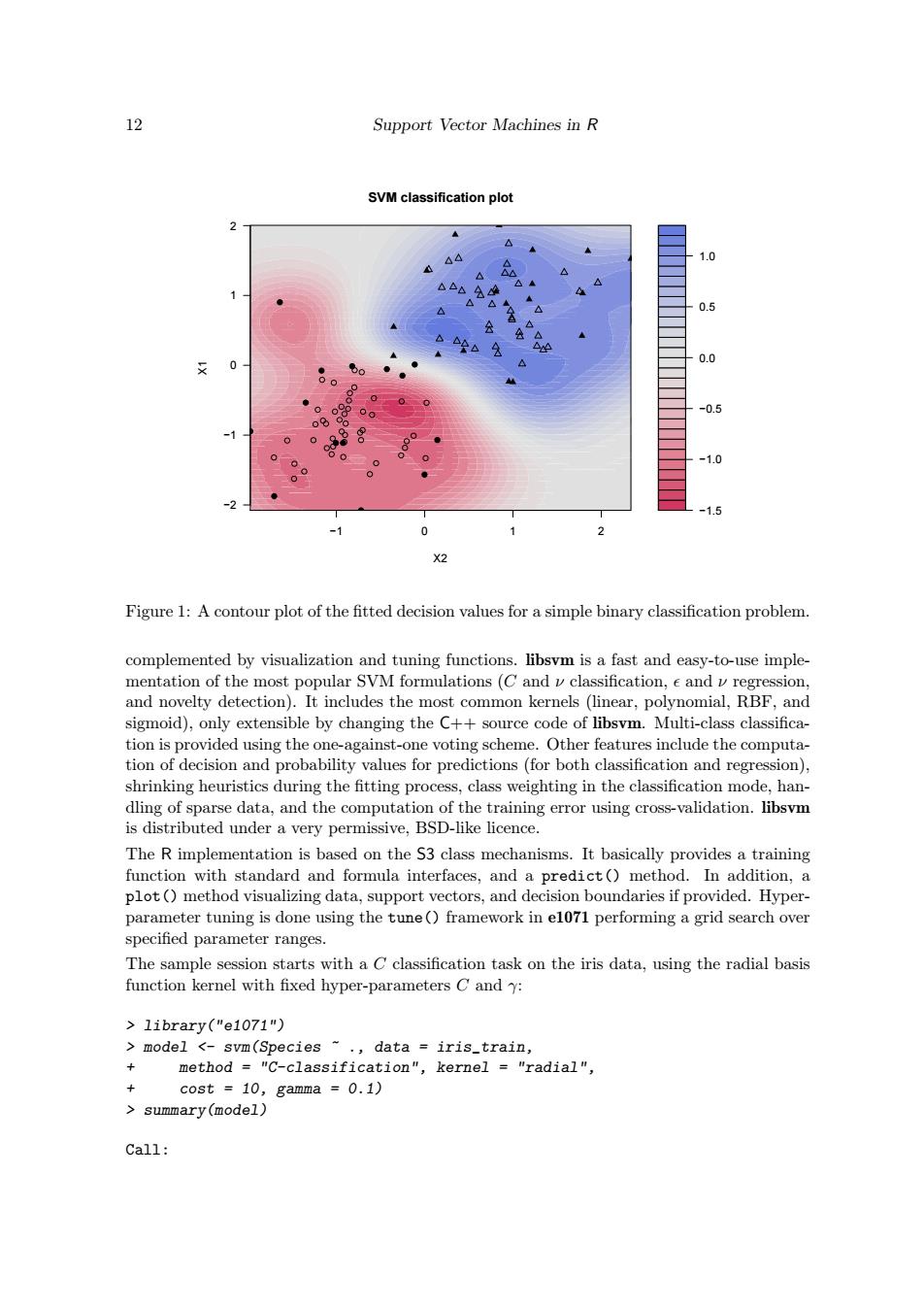
12 Support Vector Machines in R SVM classification plot 2 44 1.0 △△ △▲ 1 △△◆ 0.5 0.0 0 ● o -0.5 88 -1.0 -2 -1.5 0 X2 Figure 1:A contour plot of the fitted decision values for a simple binary classification problem. complemented by visualization and tuning functions.libsvm is a fast and easy-to-use imple- mentation of the most popular SVM formulations(C and v classification,e and v regression, and novelty detection).It includes the most common kernels (linear,polynomial,RBF,and sigmoid),only extensible by changing the C++source code of libsvm.Multi-class classifica- tion is provided using the one-against-one voting scheme.Other features include the computa- tion of decision and probability values for predictions(for both classification and regression), shrinking heuristics during the fitting process,class weighting in the classification mode,han- dling of sparse data,and the computation of the training error using cross-validation.libsvm is distributed under a very permissive,BSD-like licence. The R implementation is based on the S3 class mechanisms.It basically provides a training function with standard and formula interfaces,and a predict()method.In addition,a plot()method visualizing data,support vectors,and decision boundaries if provided.Hyper- parameter tuning is done using the tune()framework in el071 performing a grid search over specified parameter ranges. The sample session starts with a C classification task on the iris data,using the radial basis function kernel with fixed hyper-parameters C and y: library("e1071") model <-svm(Species ~.data iris_train, + method "C-classification",kernel "radial", cost =10,gamma =0.1) summary(model) Call:12 Support Vector Machines in R −1.5 −1.0 −0.5 0.0 0.5 1.0 −1 0 1 2 −2 −1 0 1 2 ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● SVM classification plot X2 X1 Figure 1: A contour plot of the fitted decision values for a simple binary classification problem. complemented by visualization and tuning functions. libsvm is a fast and easy-to-use implementation of the most popular SVM formulations (C and ν classification, and ν regression, and novelty detection). It includes the most common kernels (linear, polynomial, RBF, and sigmoid), only extensible by changing the C++ source code of libsvm. Multi-class classification is provided using the one-against-one voting scheme. Other features include the computation of decision and probability values for predictions (for both classification and regression), shrinking heuristics during the fitting process, class weighting in the classification mode, handling of sparse data, and the computation of the training error using cross-validation. libsvm is distributed under a very permissive, BSD-like licence. The R implementation is based on the S3 class mechanisms. It basically provides a training function with standard and formula interfaces, and a predict() method. In addition, a plot() method visualizing data, support vectors, and decision boundaries if provided. Hyperparameter tuning is done using the tune() framework in e1071 performing a grid search over specified parameter ranges. The sample session starts with a C classification task on the iris data, using the radial basis function kernel with fixed hyper-parameters C and γ: > library("e1071") > model <- svm(Species ~ ., data = iris_train, + method = "C-classification", kernel = "radial", + cost = 10, gamma = 0.1) > summary(model) Call: